|
2/15/2024 0 Comments Display alignment 
IGV loads this coverage track automatically when test.bam is loaded. 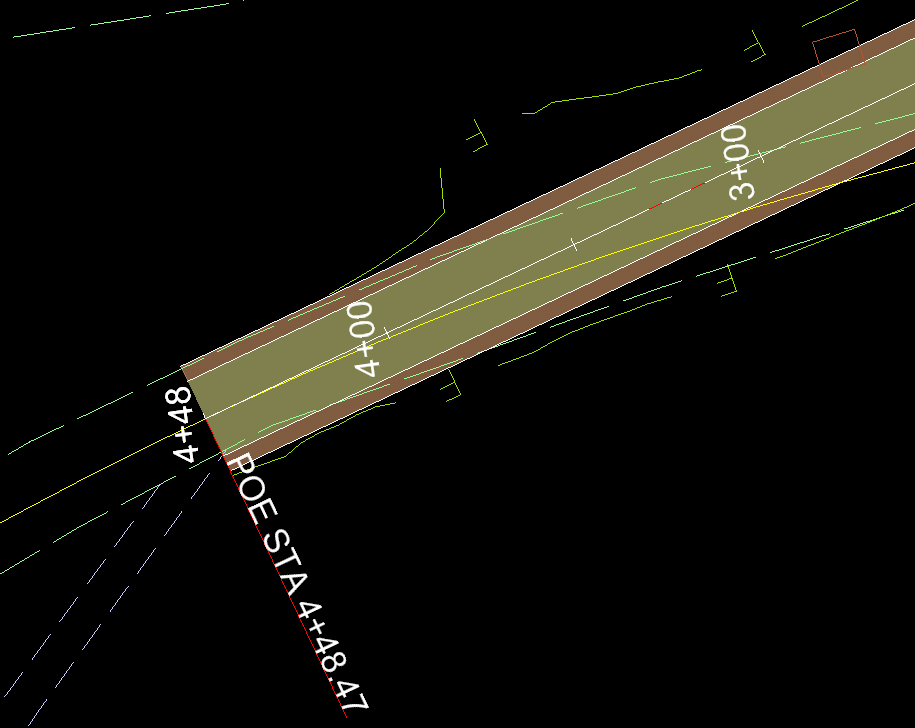
If you need to create the index yourself, there are multiple tools available for indexing BAM files, including igvtools, the samtools package, and the Picard.SortSam module in GenePattern.įor example, the coverage track for test.bam would be named. bam file from a sequencing facility, you will usually also get the corresponding index file. When loading by file, IGV automatically searches for the index file within the same directory as the data file. When loading by URL, the URL to both the data file and the index file should be specified. For example, the index file for test-xyz.bam would be named, or alternatively test-xyz.bai. The index file should have the same filename but with the. IGV requires that BAM and CRAM files have an associated index file. Related topics on other pages cover more detailed topics:Īligned reads from sequencing can be loaded into IGV in the BAM format, SAM format, or CRAM format.īoth BAM and SAM files are described on the Samtools project page and in the 2014 article titled Sequence Alignment/Map Format Specification by the SAM/BAM Format Specification Working Group. This page introduces viewing alignment data and associated tracks in the following sections:
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
